ABSTRACT
Computational models for predicting aqueous solubility from the molecular structure represent a promising strategy from the perspective of drug design and discovery. Since the first “Solubility Challenge”, these initiatives have marked the state-of-art of the modelling algorithms used to predict drug solubility. In this regard, the quality of the input experimental data and its influence on model performance has been frequently discussed. In our previous study, we developed a computational model for aqueous solubility based on recursive random forest approaches. The aim of the current commentary is to analyse the performance of this already trained predictive model on the molecules of the second “Solubility Challenge”. Even when our training set has inconsistencies related to the pH, solid form and temperature conditions of the solubility measurements, the model was able to predict the two sets from the second “Solubility Challenge” with statistics comparable to those of the top ranked models. Finally, we provided a KNIME automated workflow to predict the aqueous solubility of new drug candidates, during the early stages of drug discovery and development, for ensuring the applicability and reproducibility of our model.
KEYWORDS
Quantitative Structure-Property Relationship (QSPR); KNIME; aqueous solubility; ADME; machine learning; Random Forest; supervised recursive selection.
AUTHORS
Gabriela Falcón-Cano†, Christophe Molina∥, and Miguel Ángel Cabrera-Pérez †, §, ‡
†Unit of Modeling and Experimental Biopharmaceutics. Centro de Bioactivos Químicos. Universidad Central “Marta Abreu” de las Villas. Santa Clara 54830, Villa Clara, Cuba
∥PIKAÏROS S.A, 31650 Saint Orens de Gameville, France
§Department of Pharmacy and Pharmaceutical Technology, University of Valencia, Burjassot 46100, Valencia, Spain
‡Department of Engineering, Area of Pharmacy and Pharmaceutical Technology, Miguel Hernández
University, 03550 Sant Joan d’Alacant, Alicante, Spain
JOURNAL
DMPK & ADMET Journal (DOI: https://doi.org/10.5599/admet.979 )
ADME prediction with KNIME: A retrospective contribution to the second “Solubility Challenge”, Advance Online Articles (2021). Published: 12-07-2021
SUPPORTING INFORMATION
How to install and start with KNIME : https://www.knime.com/knime
KNIME WORKFLOW : Available at https://pikairos.eu/download/aqueous_solubility_prediction (270 Mbytes)
Please get in touch with us through our Contact Page if you have any questions concerning the journal article or the KNIME Workflow
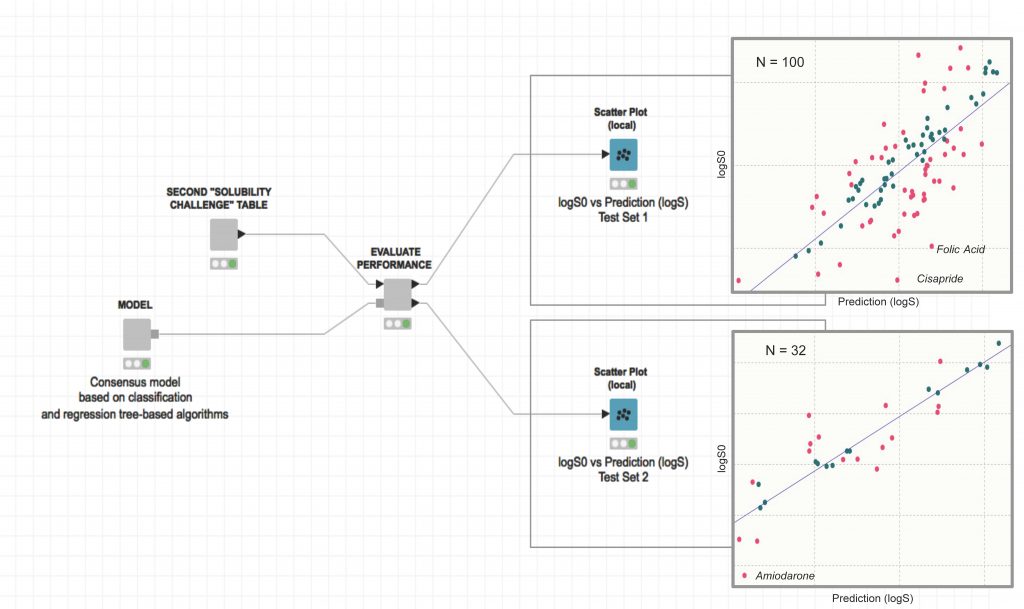
Workflow Available under the Revised BSD License
Copyright (c) 2021, Gabriela Falcón-Cano, Christophe Molina, Miguel Ángel Cabrera-Pérez & Pikaïros.
All rights reserved. Redistribution and use in source and binary forms of this workflow, with or without modification, are permitted provided that the following conditions are met:
- Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer.
- Redistributions in binary form must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution.
- Neither the name of Pikaïros nor the names of its contributors may be used to endorse or promote products derived from this software without specific prior written permission.
THIS WORKFLOW IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS “AS IS” AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE AUTHORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
